Synth4Bench
Framework for generating synthetic genomics datasets for benchmarking tumor-only somatic variant callers.
Overview
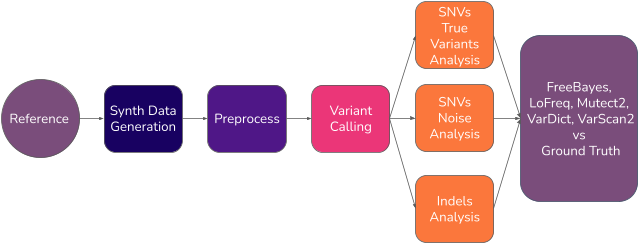
Synth4Bench is a framework for generating synthetic genomics datasets designed to support the systematic benchmarking of tumor-only somatic variant calling algorithms.
The project addresses a key challenge in cancer genomics: evaluating variant callers when high-quality ground truth data are limited or unavailable. By generating synthetic datasets with known variants, Synth4Bench enables controlled experiments where sequencing depth, read length, allele frequency and variant characteristics can be adjusted and evaluated.
My work focused on the design and implementation of the synthetic data generation workflow, the benchmarking strategy and the downstream analysis used to compare variant calling tools against known ground truth.
Code and Data
The code is available on GitHub and all data on Zenodo
Related publication
- Synth4Bench: Generating Synthetic Data for Benchmarking Tumor-Only Somatic Variant Calling Algorithms
Manuscript in preparation / under review but preprint is available in bioRxiv
Funding
This work is related to the SYNTHIA project and was developed as part of my PhD research.